Notebook
Query using the universal results warehouse table
The Benchling warehouse now has a universal results warehouse table which stores result level information in its own table: result$raw. This universal table is analogous to the existing registry_entity warehouse table. Access this warehouse table from the developer platform or through Insights queries.
Upload Box files via the external data tool
Benchling now supports an external file integration with Box. Use the external data tool to connect your Box account, browse Box files directly in Benchling, and drag and drop files directly into Notebook entries.
Note: The new Box integration will be enabled for enterprise users September 14, 2020. If you would like this new feature enabled on your tenant, please contact support@benchling.com.
Animated previews of GIFs attached to a Notebook entry
Previously animated GIFs that were uploaded to a Notebook entry rendered a thumbnail that is not animated.
Note: Entries exported as a PDF will have static GIFs.
Molecular Biology
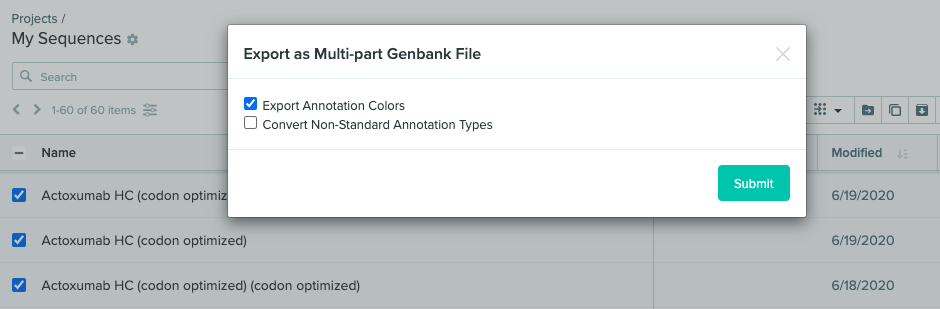
Export annotations with multi-part Genbank file exports
Benchling now supports exporting or converting annotations when exporting multi-part Genbank files in bulk. Previously, the options to “Export Annotation Colors” and “Convert Non-Standard Annotation Types” were only available when exporting a single sequence to .genbank format.
Note: If the option to convert non-standard annotation types is chosen, annotation types will map to one of the standard feature types. If an annotation type cannot be mapped to a standard feature type, the annotation will be exported as a ‘misc_feature’.
New codon optimization parameter: ‘Patterns to Reduce’
Users can now enter in custom nucleotide patterns in the codon optimization tool to reduce usage of patterns throughout the entire sequence length or the coding strand only. A checkbox also allows the user to reduce usage of the reverse complement. Note: The pattern must consist of ATGC bases only and is case-insensitive. A common use case may be to reduce the presence of CG motifs.
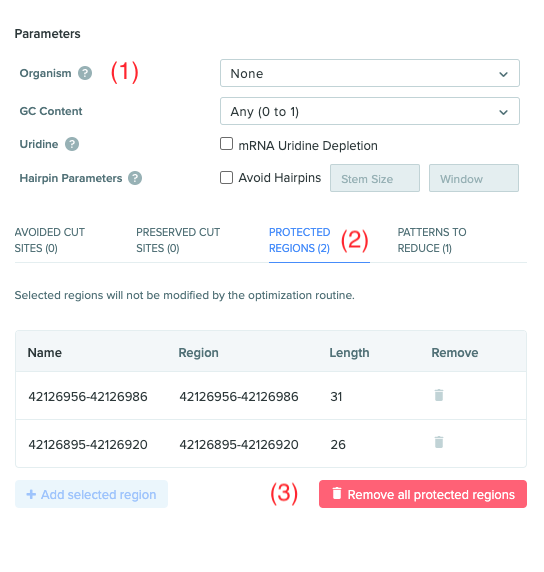
User interface polishes to codon optimization
(1) Organism is now an optional parameter so that other parameters can be properly used, (e.g. ‘Patterns to Reduce’) without having a competing constraint. (2) Each of the avoided cut sites, protected regions, etc are counted and numbered to highlight how many constraints of each type are configured. (3) “Remove all” button makes it easier to clear out existing constraints. This button is visible for “Avoided Cutsites”, “Preserved Cutsites”, “Protected Regions”, and “Patterns to Reduce”
Registry and Inventory
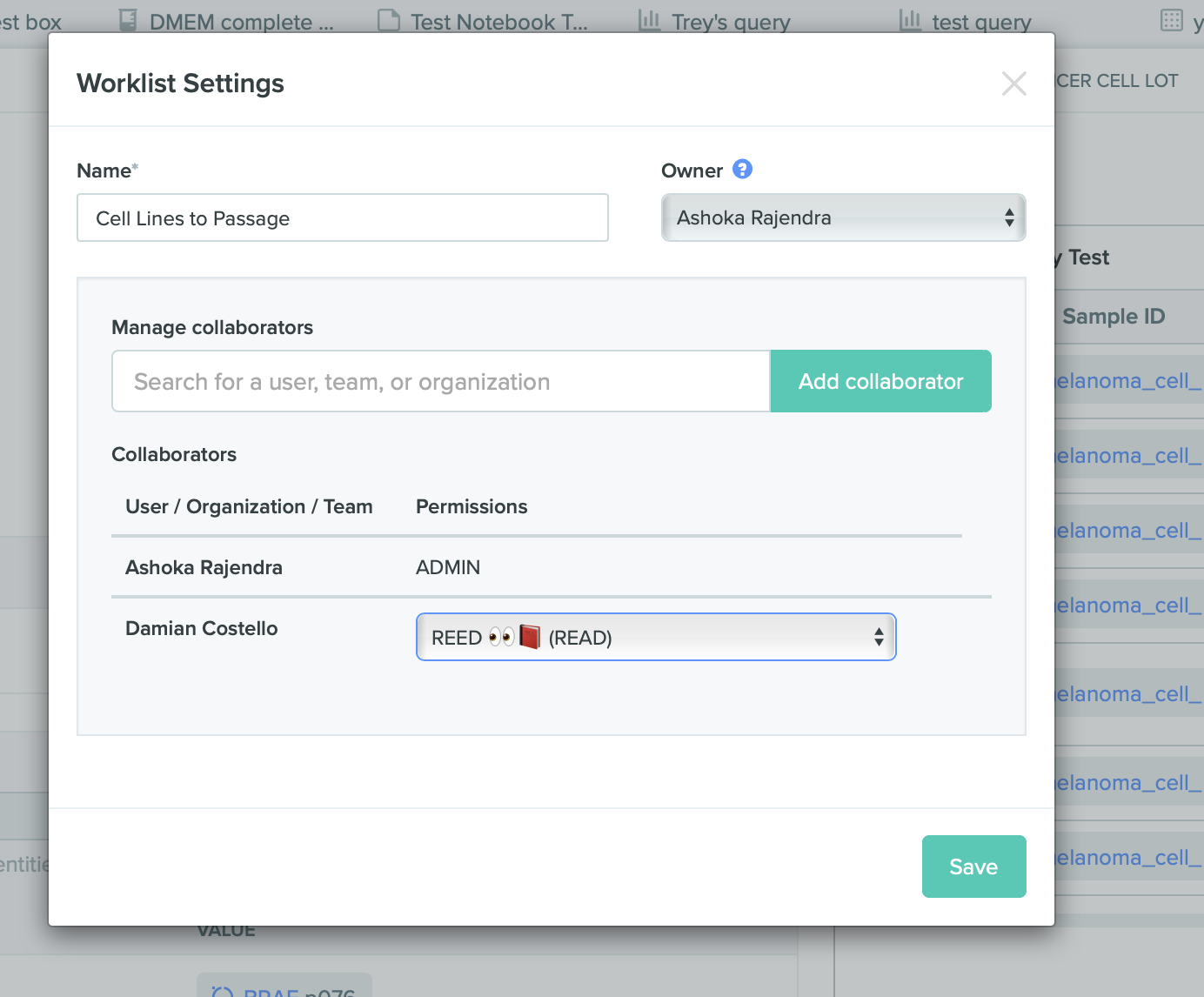
Share worklists with users, teams, or orgs
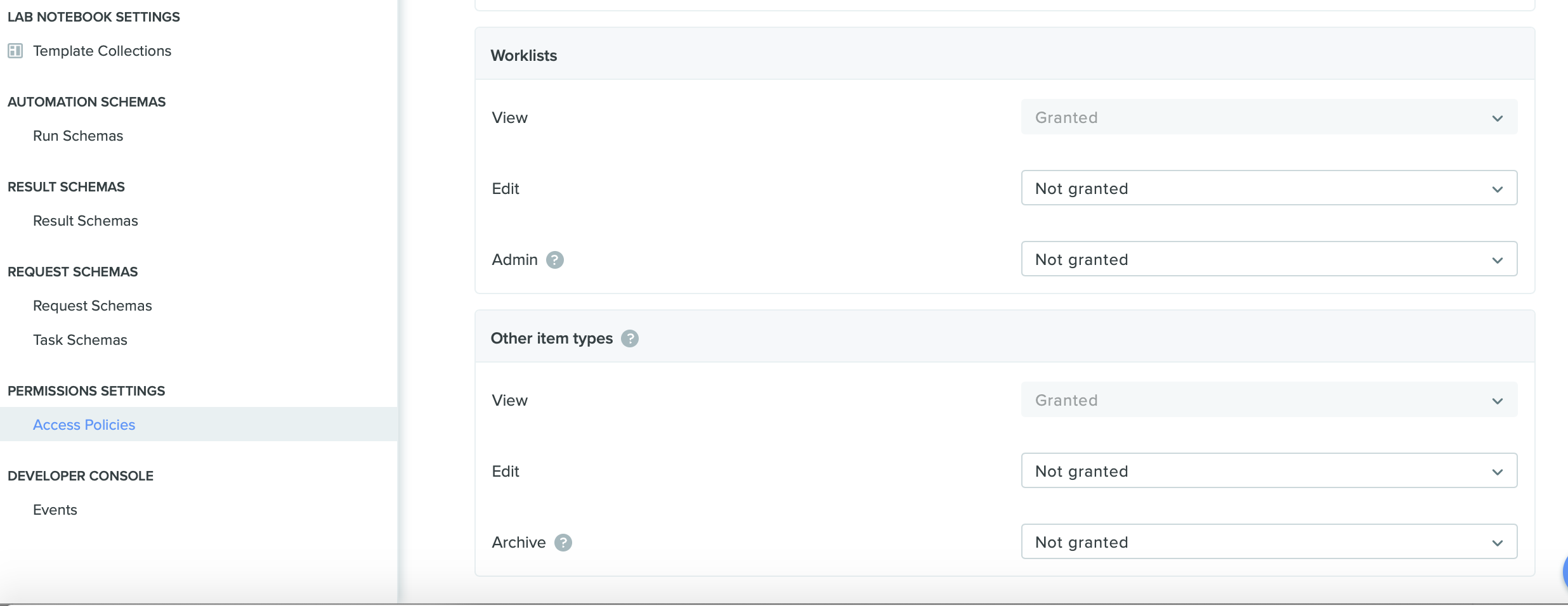
Benchling now supports the option to share worklists with other users, teams, or orgs. Previously, worklists could only be used and viewed by the creator of the worklist. However, users can now share worklists by clicking into a worklist and accessing the worklist settings. Additionally, control worklist permissions using access policies to grant the ability to view, edit, or allow admin rights to a worklist.
Delete a contained item from a container via the API
Our developer platform now supports the option to delete a contained item from a container. To explore this new API endpoint further, please review our API resources here.
Receive concise Registry validation emails
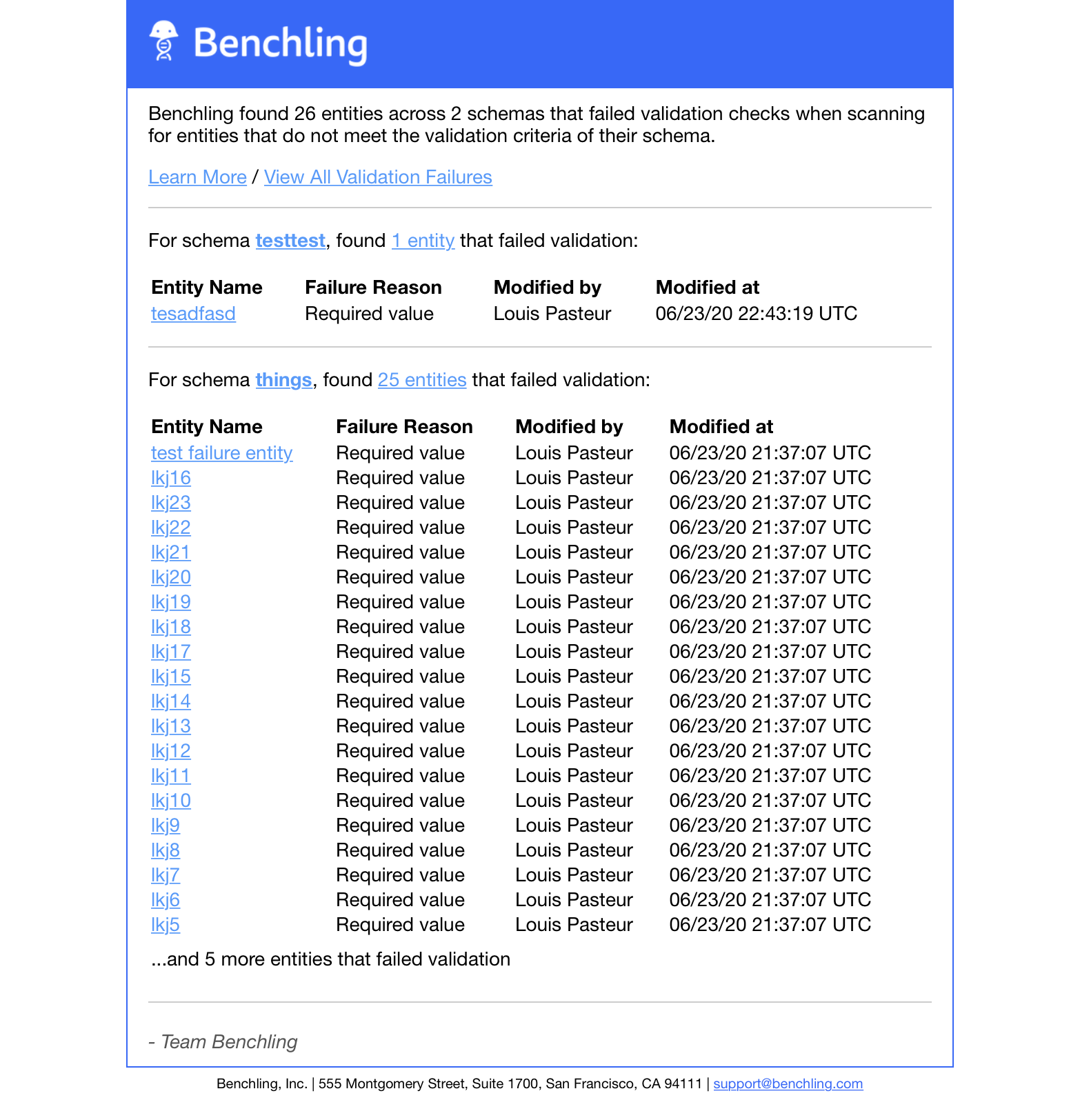
Previously, emails describing validation errors would be sent to Registry admins as soon as an entity failed validation. However, to avoid excess emails and alert fatigue, Registry validation errors are now summarized and aggregated into a single email sent once per day. If there are no Registry validation errors, no email will be sent to Registry admins.
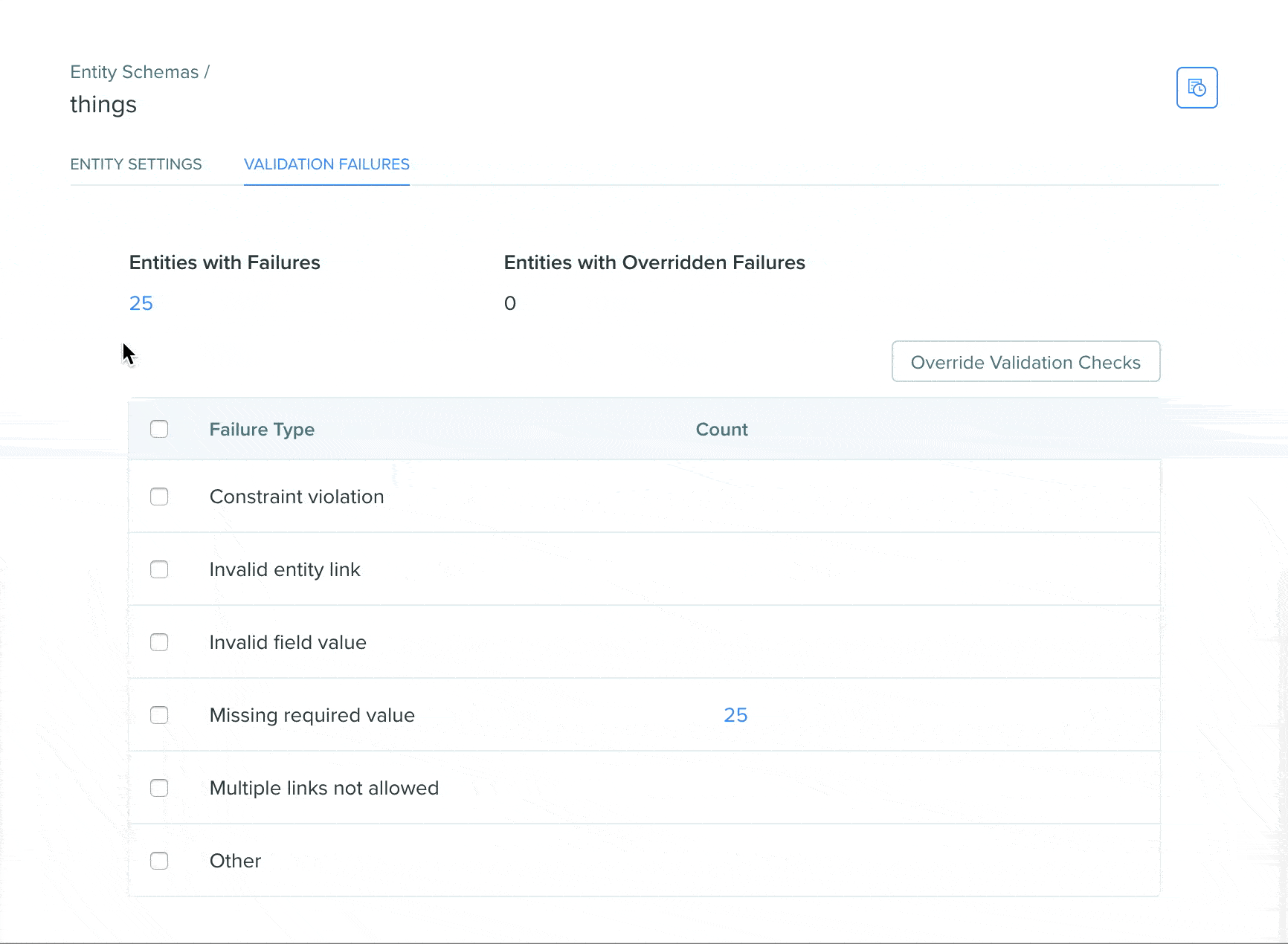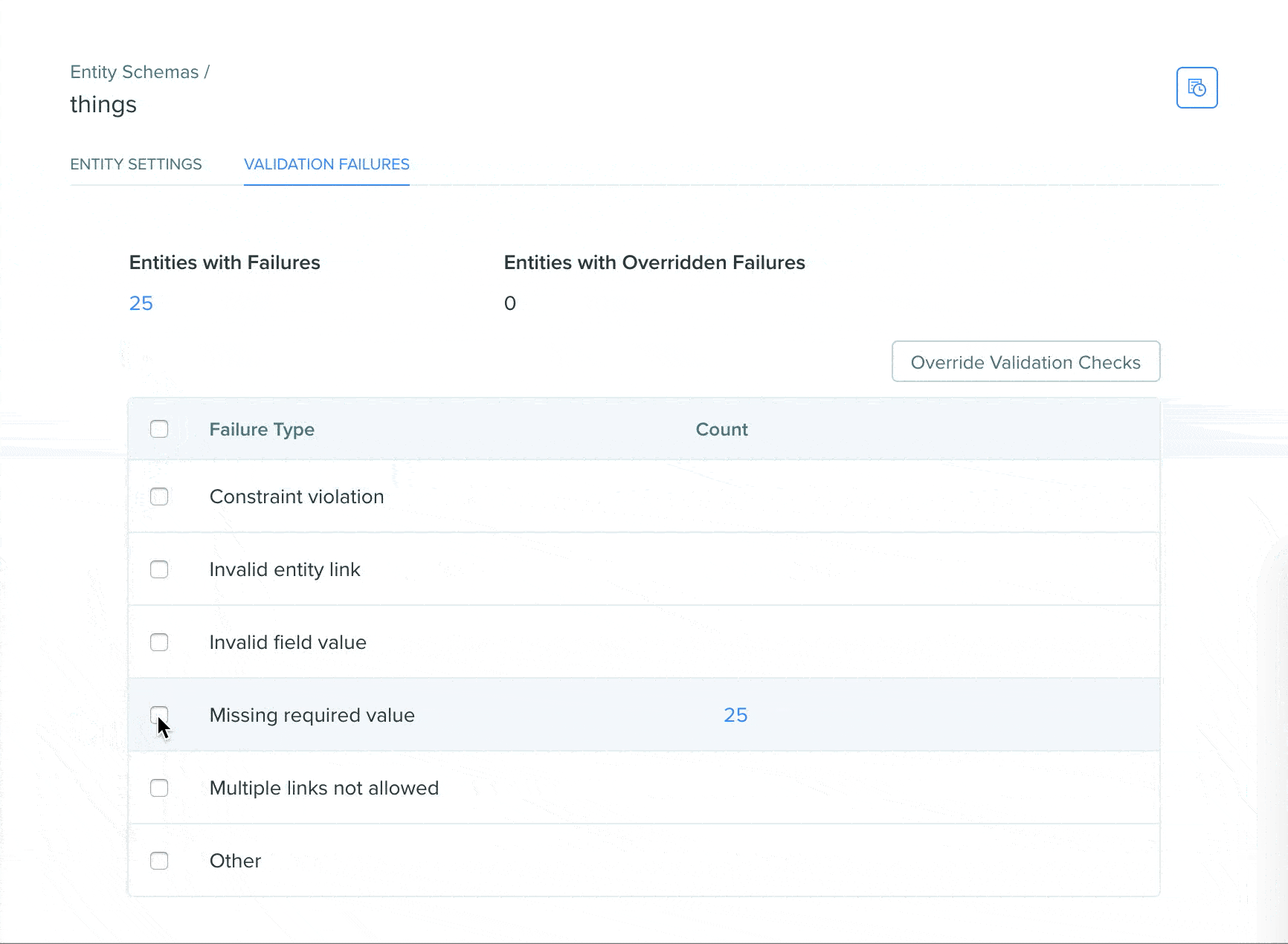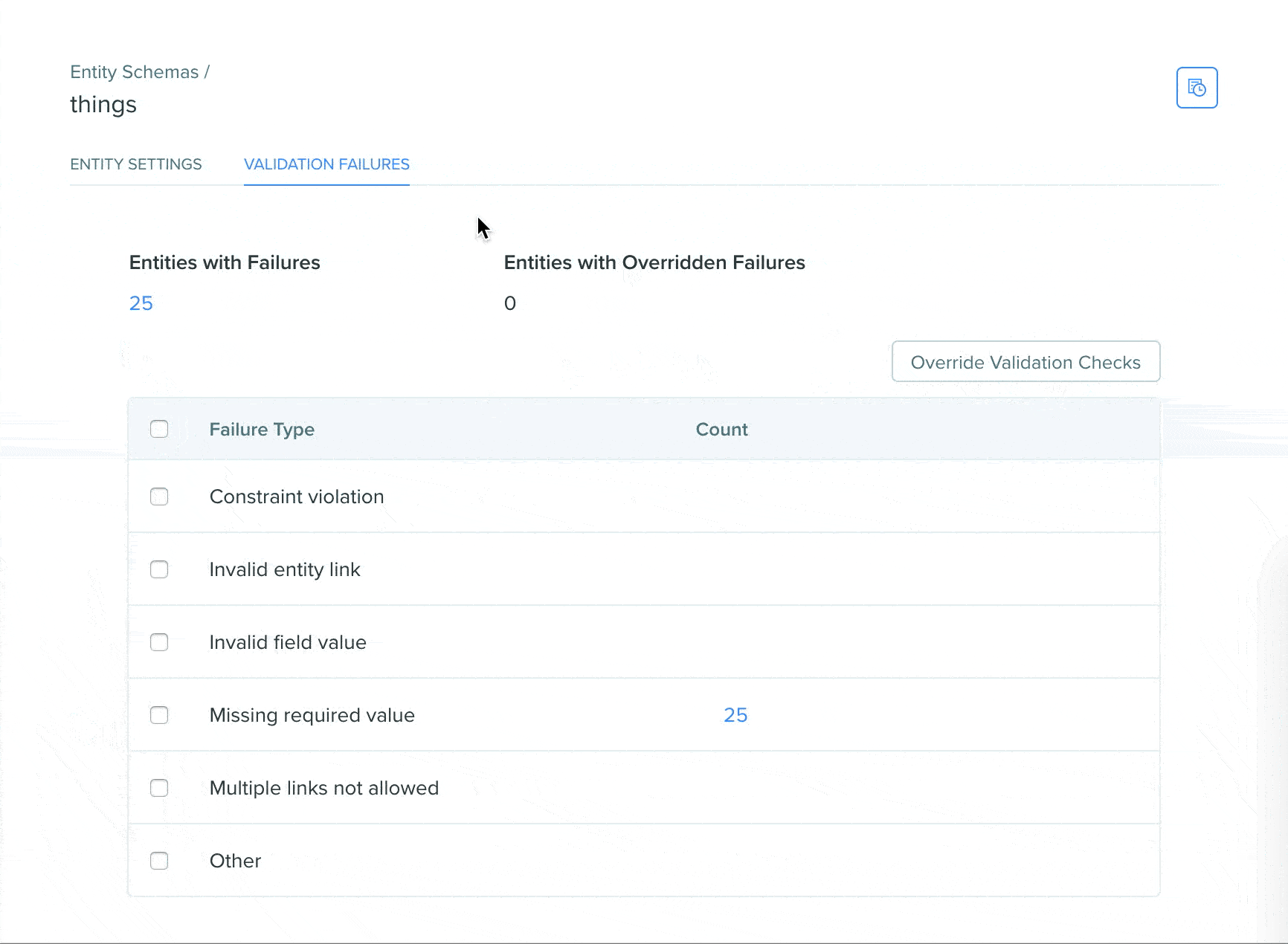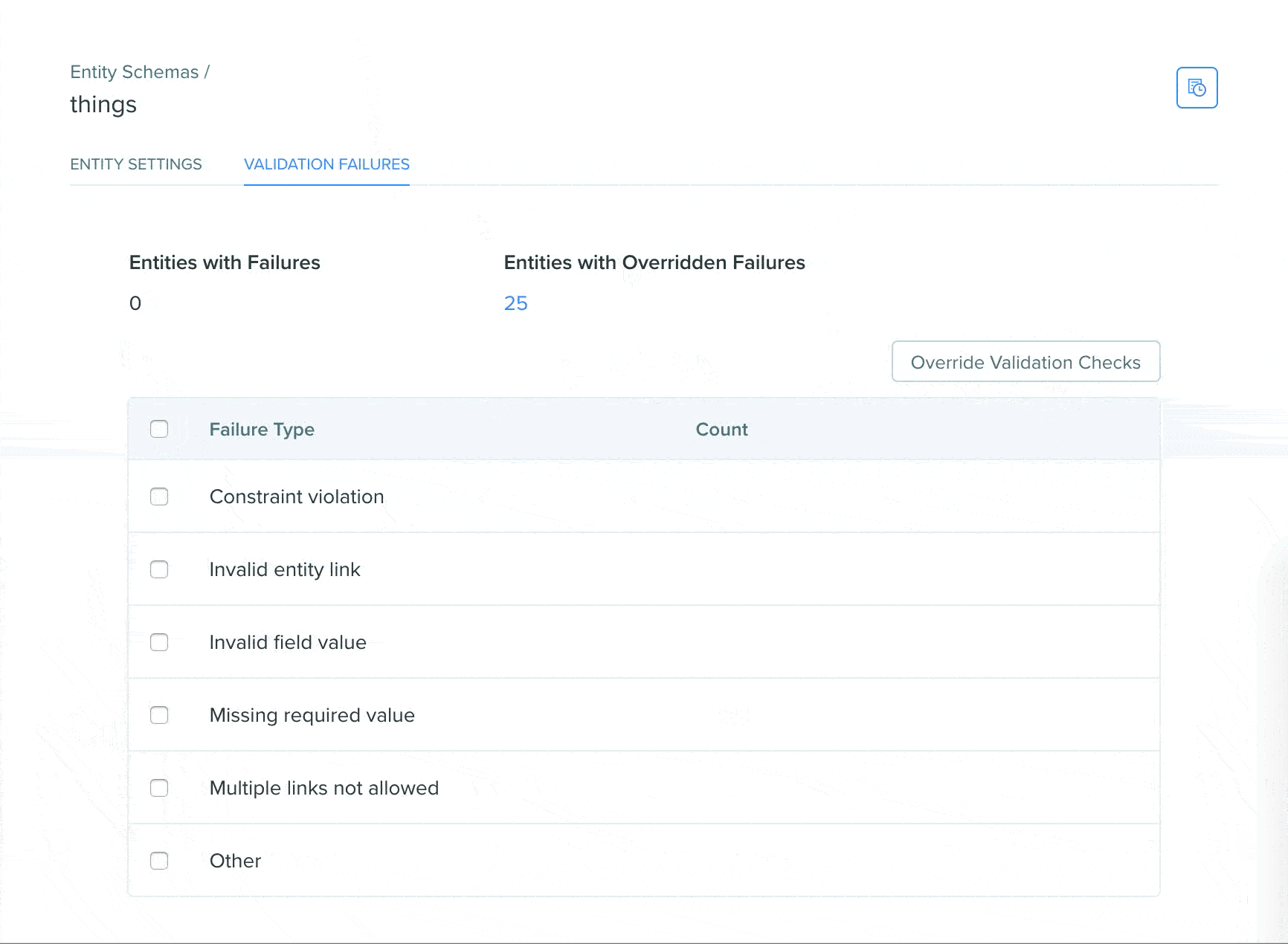
View validation status using the Registry validation tab
A Registry validation tab has been added to the Registry schema page of DNA, AA, and custom entity schemas. The validation tab allows users to easily view validation failures in a summarized format on the Registry schema page, click into failures to see the entities in a search view, and easily bulk override failures.
Mixture preparation tables in the Notebook
Mixtures are solutions comprised of multiple ingredients where the exact quantities of each ingredient are important to track as they can be set as a template for preps. The new mixture preparation table now natively supports prep creation, not just in the Registry, but now in the Notebook, as well. Users specify the recipe and record measured amounts and lot numbers. If you would like this new feature enabled on your tenant, please contact support@benchling.com.
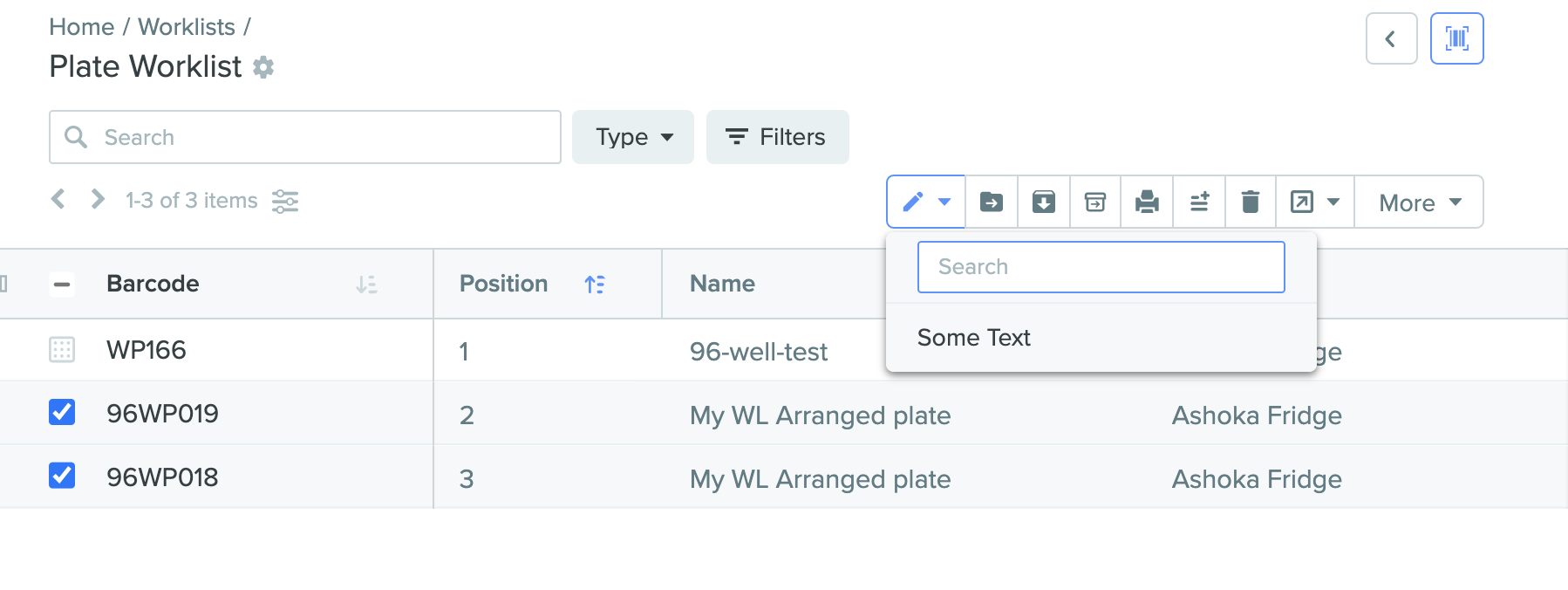
Bulk set plate fields in plate worklist
Plate fields can now be set in bulk in the expanded view of a worklist, similar to entities and plates. This action can be found by selecting the first icon in the toolbar above the worklist.
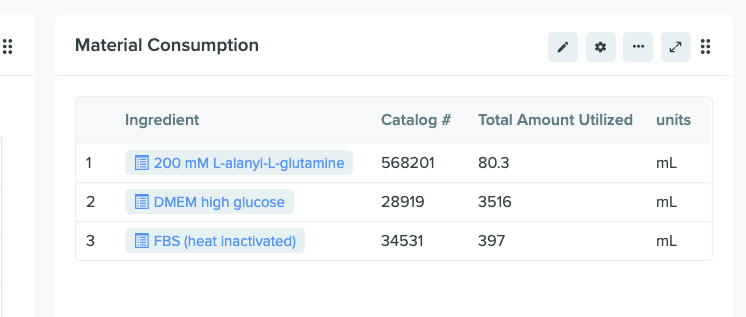
Mixture ingredient data is in the data warehouse
Look for the new mixture ingredients in the warehouse by searching for <tenant>.mixture_ingredient. The warehouse table will house mass and volume information, units, in addition to other metadata for each ingredient. If you would like this new feature enabled on your tenant, please contact support@benchling.com.
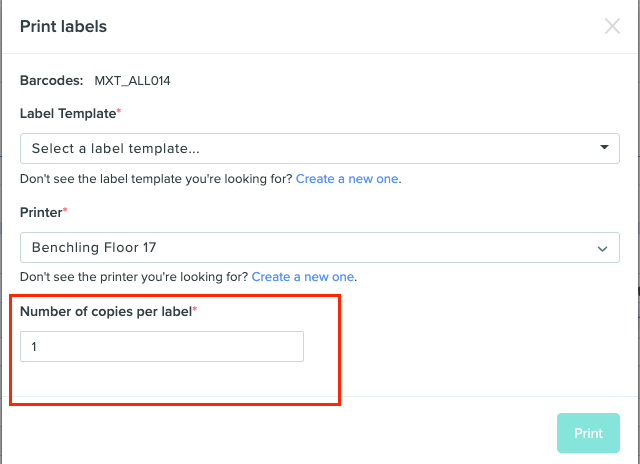
Print multiple copies per label
Previously users lacked the ability to print the same label multiple times for 1 object, unless done manually. This feature has added a field in the print labels modal that specifies the number of copies per label to be printed.
Schemas
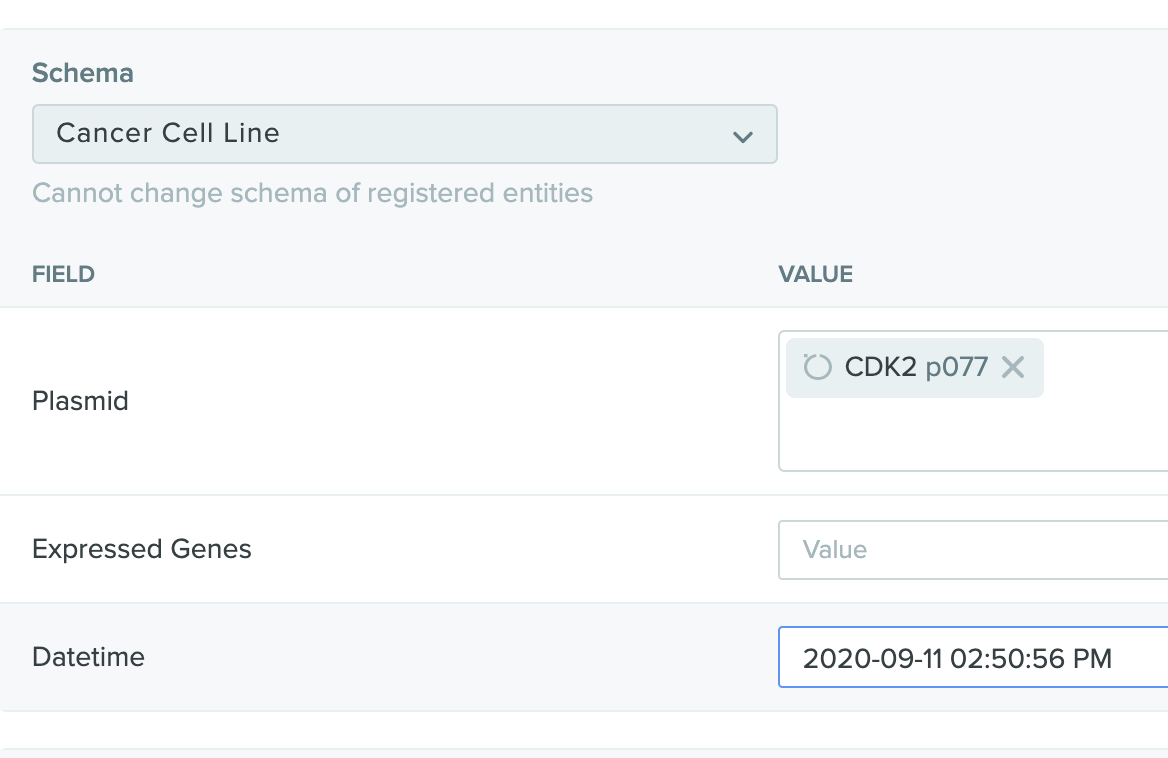
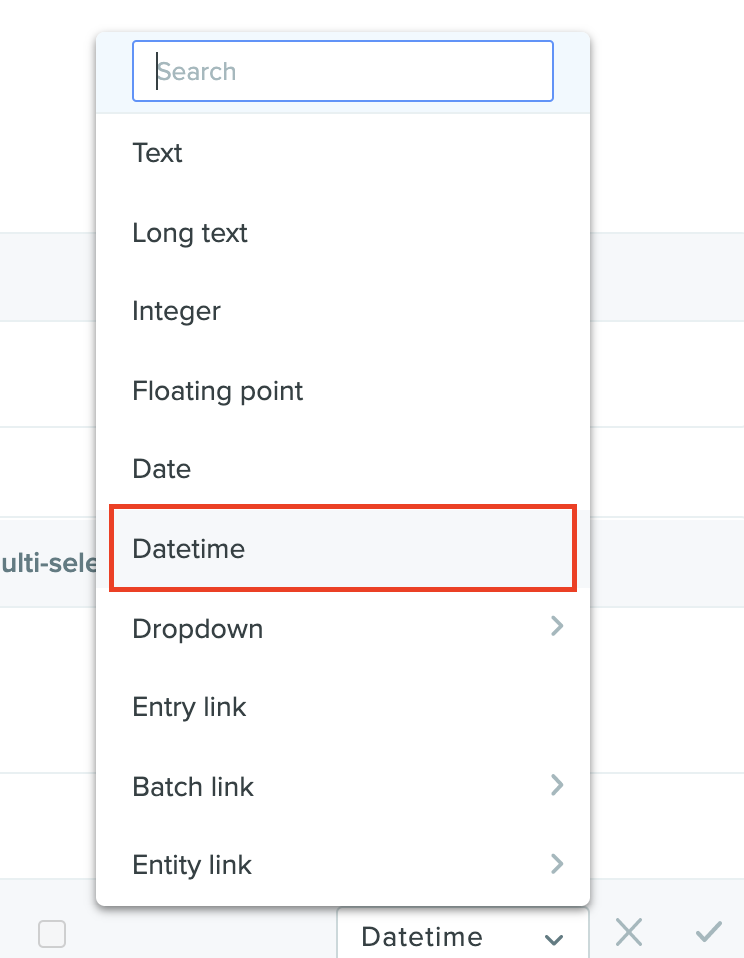
Registry schemas support datetime as a field type
Schema fields now support datetime fields in the following format: YYYY-MM-DD hh:mm:ss AM/PM. Entities can be updated with this new field either via the UI or through Benchling’s API. Create, update, and visualize entities’ datetime field values in the metadata tab view, entity create modal, or entity registration modal. If you would like this new feature enabled on your tenant, please contact support@benchling.com.
Lab Automation
Remove trailing rows from output files
Configure the processing step of a run schema to remove trailing rows from output files. This is useful for trimming footers on output files. For example, the spec below could be used to trim a footer consisting of three rows.

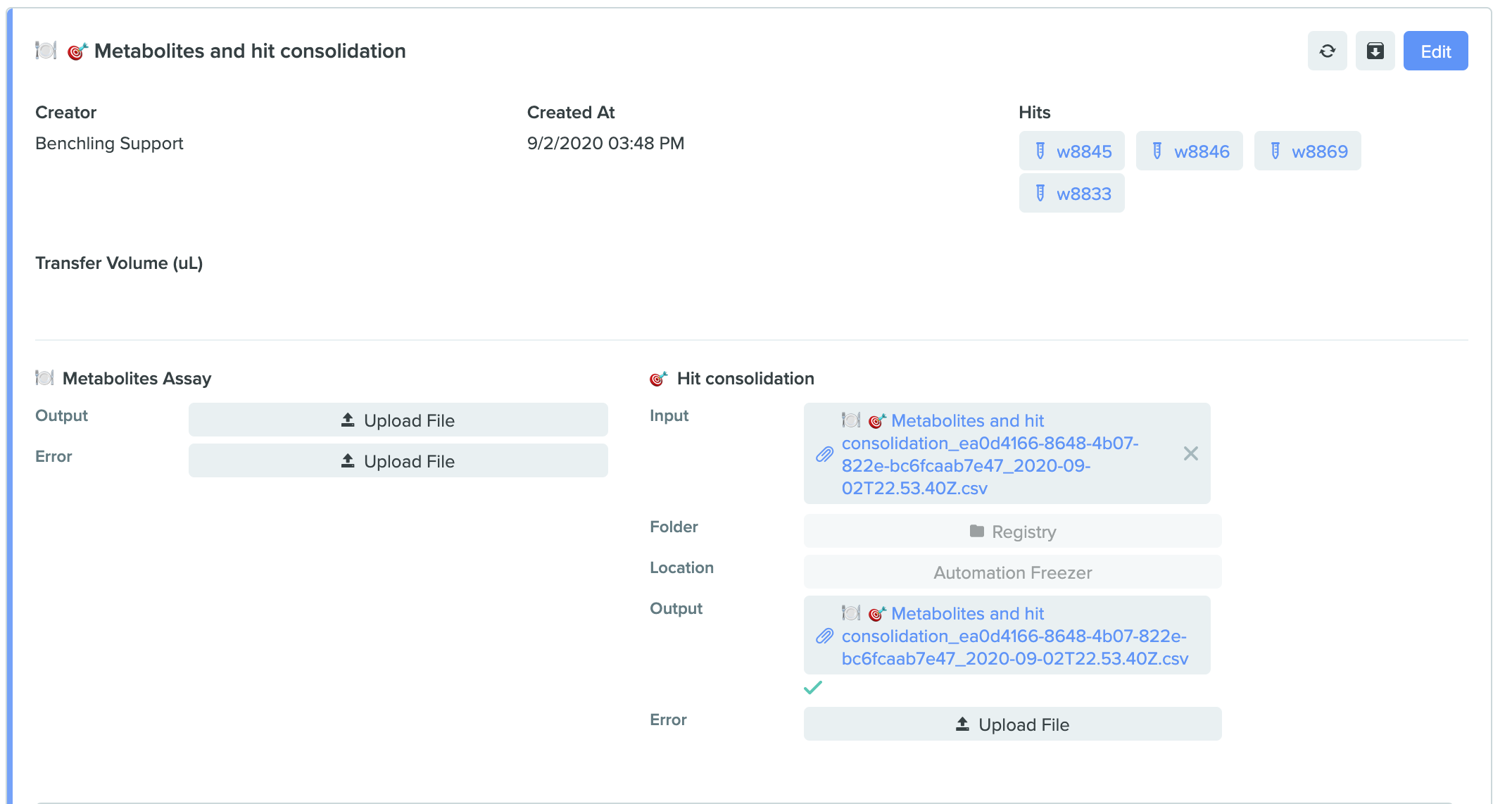
Select project folder and location in runs
Benchling now allows users to select a "Project Folder" and "Storage Location" in Lab Automation runs. The destination information for the run inputs can now be set at runtime, instead of in the run schema itself. Configure these new fields on the run schema to allow the following behavior:
-
folder: allows the user to select a folder (or the registry) for both entities and storage objects when the
projectisnull. -
location: allows the user to select a location for storage objects when the
locationisnull.
Filter out results rows with certain values
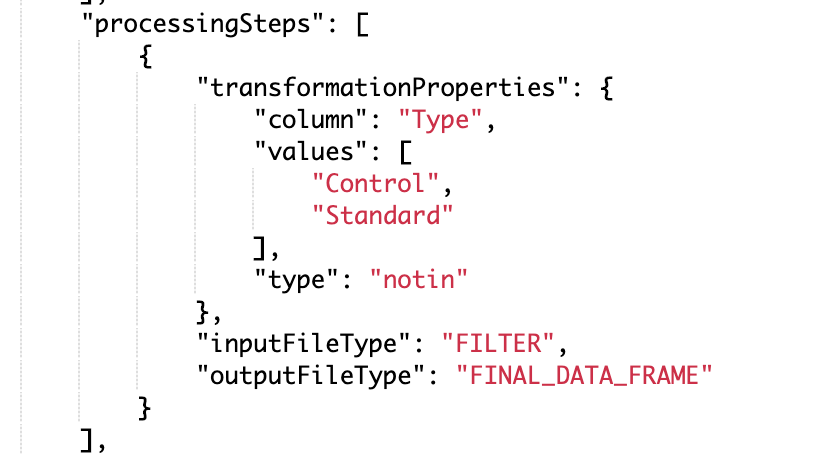
Configure a run schema with the new filter type “notin” to filter out rows that contain certain values in output files. In the example spreadsheet image below, this filter can be used to ignore standards on a plate (i.e. STD1, STD2, STD3, etc.).

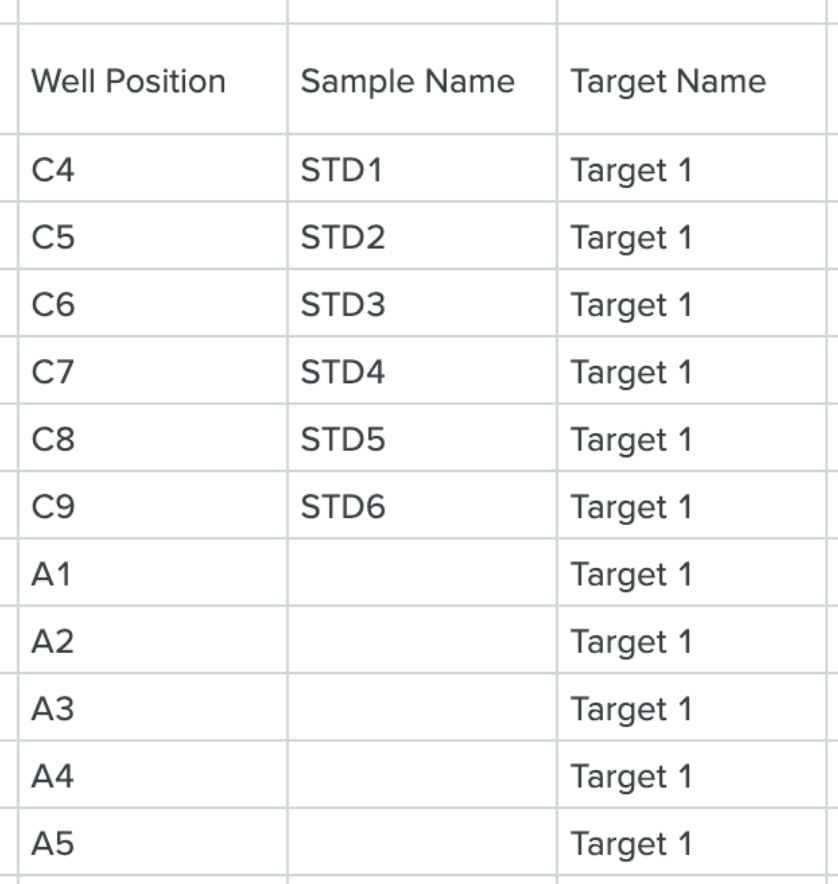
COUNT lookup step returns the count of values in a schema field
A new LOOKUP step has been added to the LookupConfig portion of the input file. This LOOKUP step counts the number of values in a schema field.

Input and output files support configurable delimiter and file extension
The input and the output file configs now support two additional properties: delimiter and file extension. Delimiter is a single character, defaulting to (and hidden if set to) “,” and controls the delimiter for parsing output files and saving processing files (if processing steps exist). fileExtension is a string defaulting to (and hidden if set to) “csv” and controls the file extension for generated output processing files (if processing steps exist).
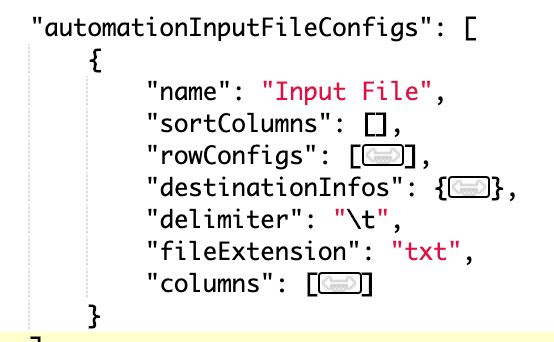
Upload results via Lab Automation where the result schema has no primary sample
Lab Automation now supports uploads into results tables via Lab Automation with files that contain null values in the results' primary entity field. This update is now consistent with normal result schemas supporting a primary sample type of none. Users will now be able to record results against control wells that aren't related to an entity.
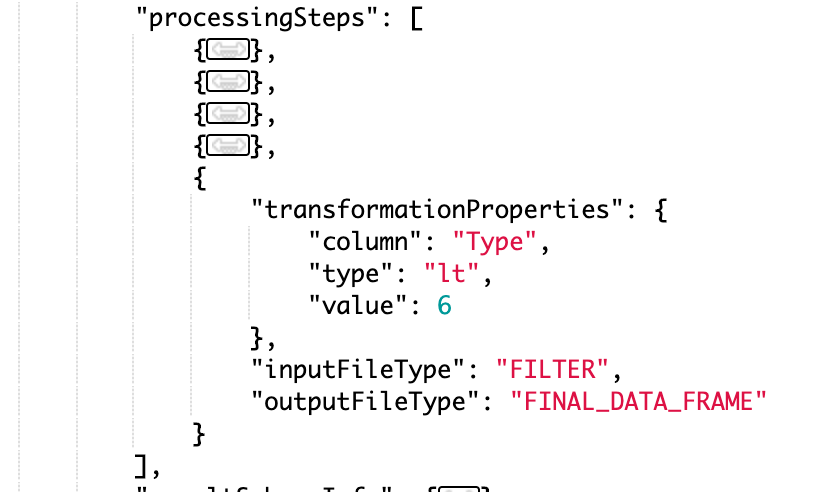
Output file filtering on numerical ranges
Filter values in the Lab Automation processing step based on numerical ranges, including less/greater than, less/greater than or equal to, equals, not equal, and between. These numeric comparisons require a numeric “value” key.

App Platform
Remove character limit restriction on handles for enterprise tenants
Enterprises with SAML authentication might have usernames in their identity provider that have one or two characters. Usernames of any length are now allowed on enterprise tenants.
Dev Platform
New endpoints to retrieve run, request, and result schemas via their unique ID
To improve coverage of Benchling’s API, new endpoints have been added for run, request, and result schemas that allow you to retrieve them by their unique ID. Please reference our API docs for more detailed information:
-
Result: https://docs.benchling.com/reference#get-assay-schema
-
Request: https://docs.benchling.com/reference#get-a-request-schema
Workflows
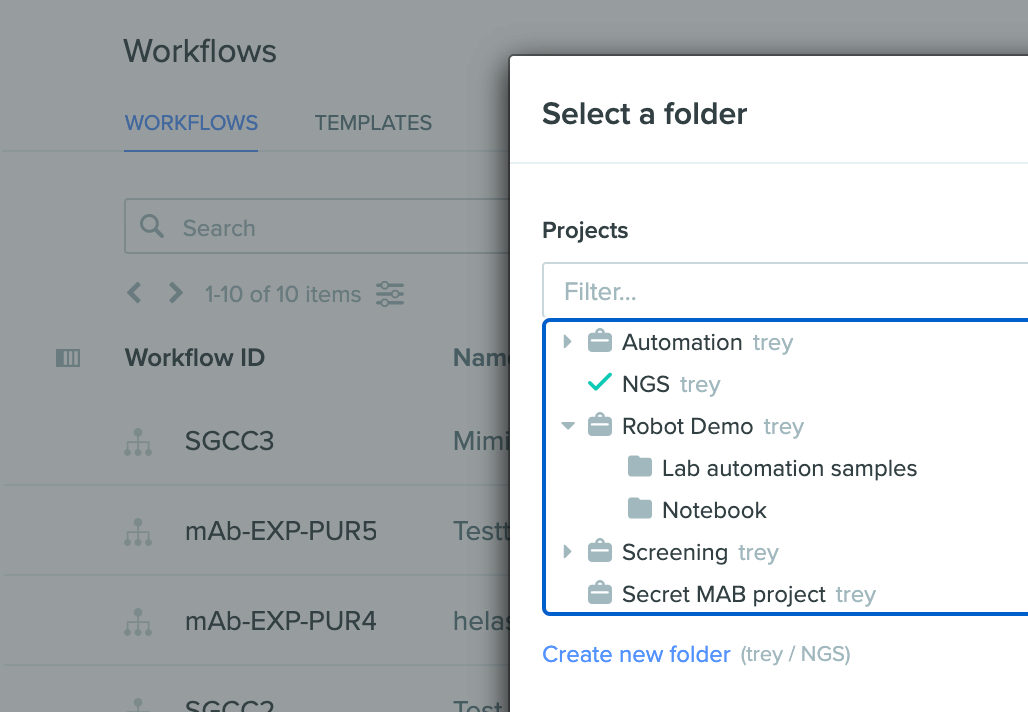
Select a folder instead of a project when creating a workflow
When creating a workflow, the create workflow modal now allows the user to select a folder as the location for the workflow instead of the project. The option to select a folder location ensures that any entities created during the workflow do not flood the root project level and instead go into this folder.
Bug Fixes
The following bugs were fixed in this release:
Dev Platform
-
Lab Automation APIs now properly create results tables in the UI.
Schemas
-
Record and display precision of trailing 0’s. Previously, trailing zeros in float fields were truncated when they were stored across the product (ie Notebook, capillary, warehouse). This precision is now retained.
Lab Automation
-
Previously, run schema processing steps that included “notnull” would not filter null rows. However, this has been fixed so the “notnull” processing step filters null rows.
-
Previously, the Request template Project ID was not respected when creating a Request from a structured table. This has been fixed so the specified project ID is respected.
-
Under certain conditions, when executing a Request all tasks would be executed despite only selecting certain tasks for execution. This has been fixed so only tasks selected for execution should appear in fulfilling Notebook entries.
-
Previously, Requests created via the API did not respect the autoAdd logic in task templates. This has been fixed so all Requests created via the API will respect autoAdd (e.g. Task templates with “autoAdd” = “false” will not automatically populate on a Request created via the API).
If you’re enjoying using Benchling, refer a friend and we’ll donate $100 to support STEM education! Learn more here