When creating translations, Benchling automatically detects and inputs valid translation links to associate amino acid entities to the DNA entities that encode them. This metadata field helps you track these relationships and ensure data integrity within your Registry.
Auto-fill translations automate the process of completing translation links by discovering registered AA sequences of the specified schema that can be validly created off one of the six reading frames of the DNA sequence you’re auto-filling.
This article explains auto-fill translations with an example and how auto-fill translations work on translations that use alternative genetic code. To configure translation links, visit Configure a translation link.
Translation link example
The image below shows the DNA sequence HC-DNA-888 with the DNA sequence schema Chain_DNA.
The image below shows the two Translation Link fields on the schema:
- Protein: a single select Translation Link field that links to the AA schema Chain”
- Multi protein: a multi select Translation Link field that links to the AA schema Recombinant Protein
Based on these schema details, the schema Chain_DNA allows us to link:
- A any single registered AA entity with the schema Chain
- Any number of registered AA entities with the schema Recombinant Protein
Note: If Benchling identifies multiple valid AA entities for a single-select Translation Link field, you might encounter an error. You can configure a Translation Link field as a multi-select field in the Registry Settings for the schema.
Benchling scans through all registered AA entities of the schema types specified in the Translation Link schema fields (Chain and Recombinant Protein), and:
- Adds translation links to all valid translated AA entities of the appropriate schema(s)
- Creates a matching translation on the sequence map to visualize the translated region
If translation links already exist on the DNA sequence you’re auto-filling translations on, the existing links are re-validated and invalid links are automatically removed.
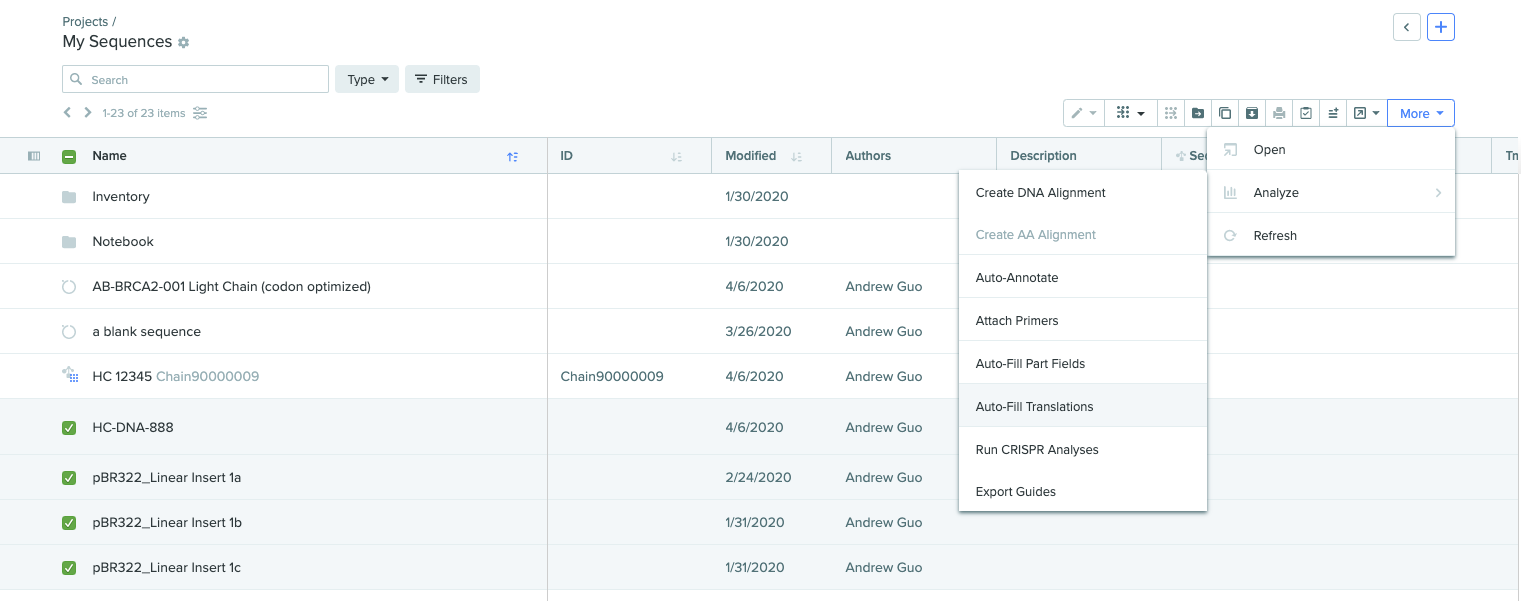
To auto-fill translations on multiple DNA sequences:
- Click > to expand the Projects panel.
- Select the sequence entities.
Click More in the top-right corner, select Analyze, then select AutoFill Translations.
Genetic codes on translations
Autofill translations use standard genetic codes to find translations. However, you can link a translation from an alternative genetic code if there is already a translation on the sequence with the same genetic code.
You can change the genetic code for a linked translation, but the translation link will become invalid.