Welcome to the April edition of our product release newsletter! We have been working hard to bring you some much requested features over the past few months. This month’s spotlight is on two feature sets. Read along to learn about how these features can help accelerate and streamline your research.
New project organization and filters
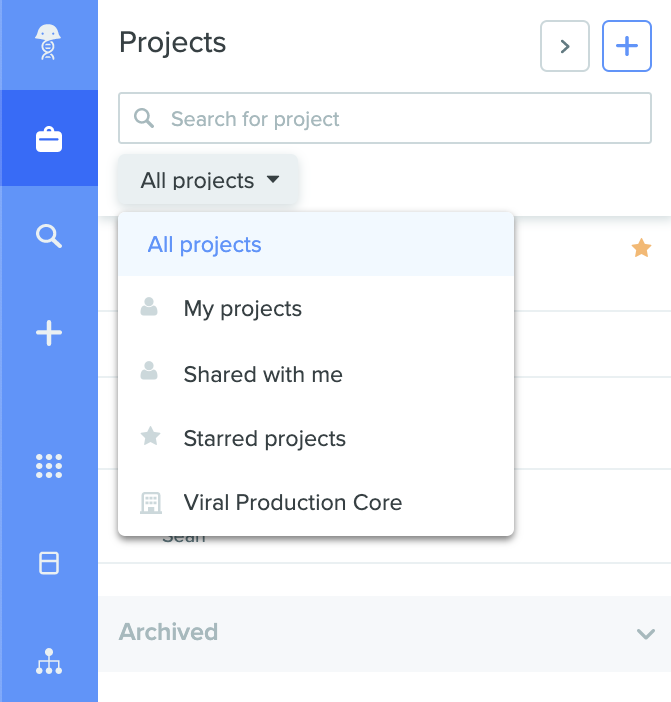
Projects are the default way in which all Notebook entries are organized, accessed, and shared on Benchling. As R&D has become more collaborative and highly complex, this has led to increasing number of experiments that need to be shared among team members. The new R&D paradigm demands a more intuitive, flexible, and practical way of collaborating across R&D organizations. We recently overhauled our project filters to meet this need. Three new project filters are now available. ‘My projects’ filters for projects with Notebook entries and Inventory items where you are the creator. ‘Shared with me’ filters for projects with Notebook entries and Inventory items that have been shared with you by other members of your team, and ‘Starred projects’ filters projects you have starred personally. Note that these filters show up only if projects with Notebook entries / Inventory items fitting these categories are available. Thus, scientists can now sort through large sets of Notebook entries and Inventory items with ease and get to the relevant ones quickly, saving them time and effort.
New APIs to unlock the power of Registry
-
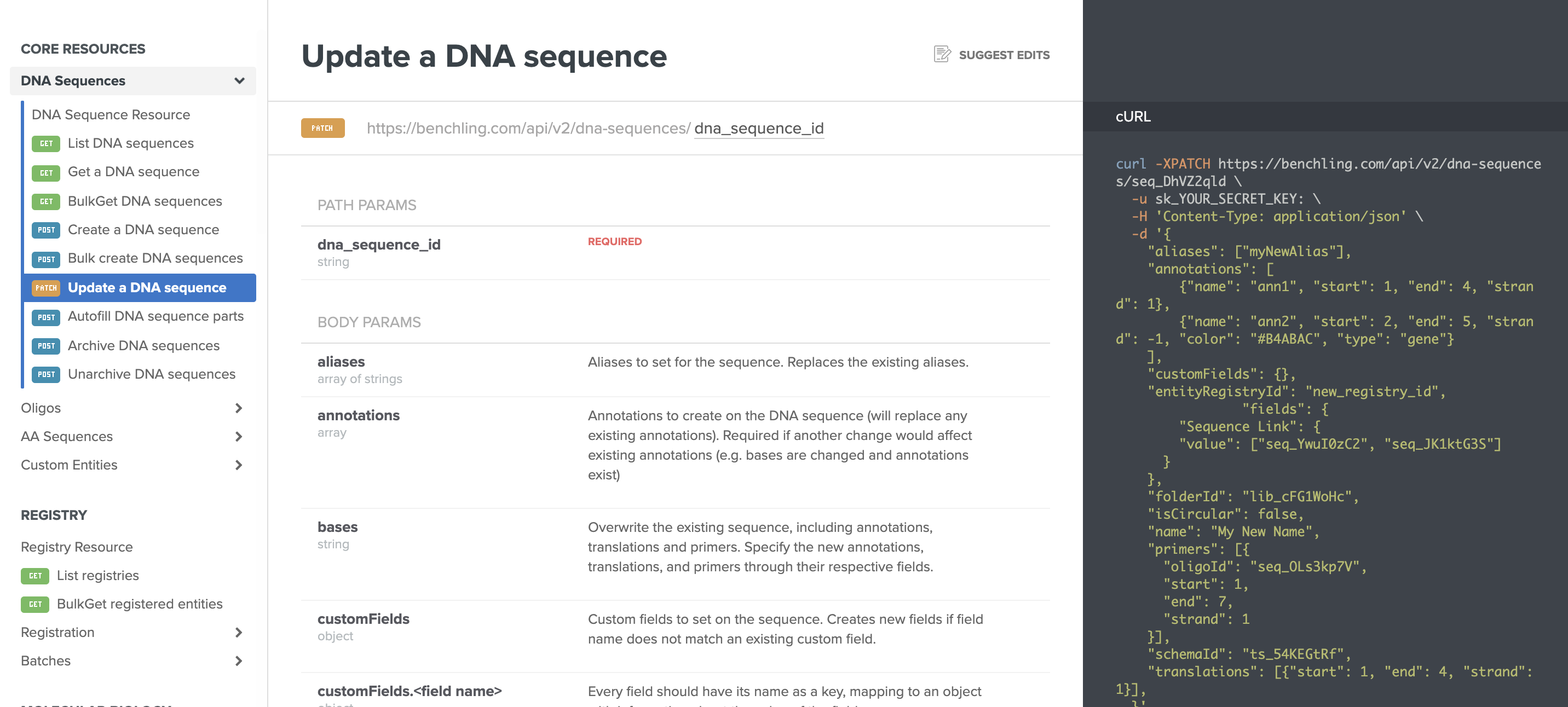
Update Registry IDs in the Benchling API: Benchling now enables you to update default Registry IDs with custom IDs through the Benchling APIs. This helps support the unique naming needs of diverse R&D programs. Learn more about this feature by visiting the links for DNA sequences, proteins, and custom entities.
-
Retry logic with bulk registration API: Previously, simultaneous registration attempts within a single Registry entry resulted in an error response. With the new API endpoint, retry logic is now built in. Up to 3 automatic re-attempts are performed before alerting the user. This helps minimize disruptions to bulk registration.
For a full list of all our major releases by product see below.
Notebook
Benchling’s Notebook was designed to make record keeping intuitive and still meet the exacting standards of today’s R&D organizations. Recent releases improve the audit log for rejected Notebook entries and expand the functionality of Benchling Tables with new powerful formulas.
-
Require comments before rejecting Notebook entries: Benchling now has the option to require comments to be recorded before Notebook entries are rejected during the sign-off process. This ensures that the full context behind this critical action is captured in the audit log for future reference. Contact support@benchling.com if you want to have this feature turned on for your team’s use.
-
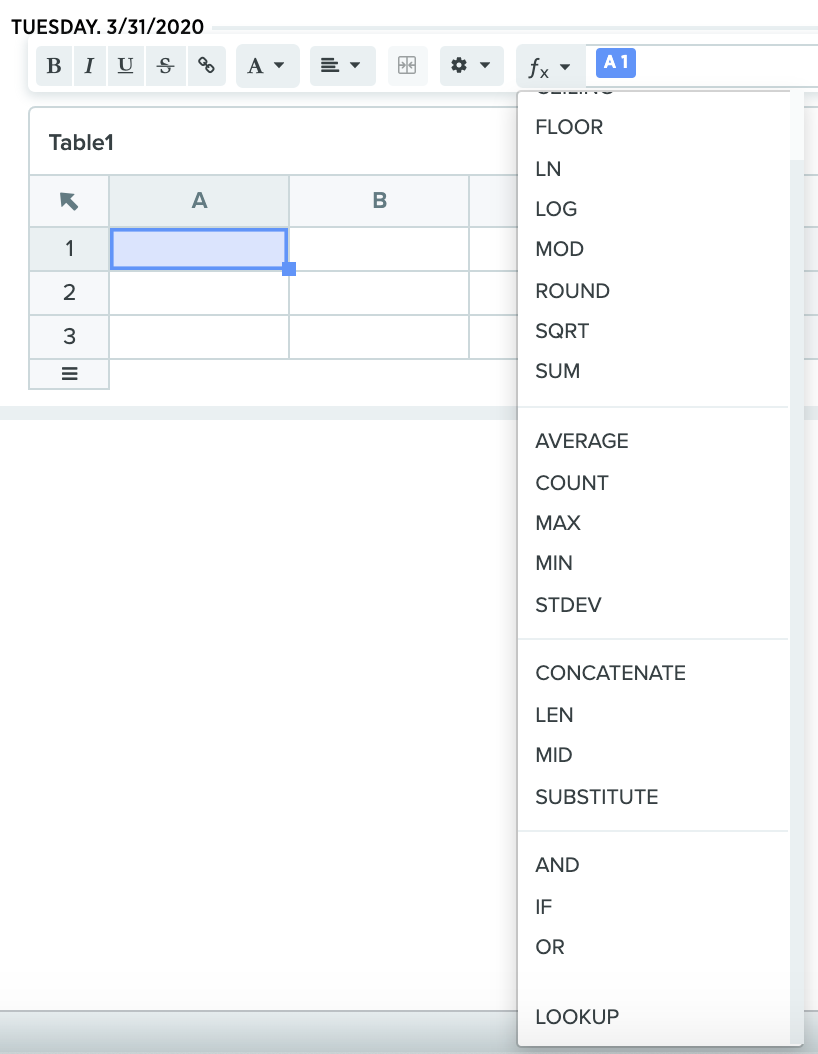
AND | OR | SUBSTITUTE functions in Tables: A few of the most popular spreadsheet functions are now available in all Benchling Tables. These functions are extremely versatile and can be used to test multiple conditions or replace content in the Table cells.
-
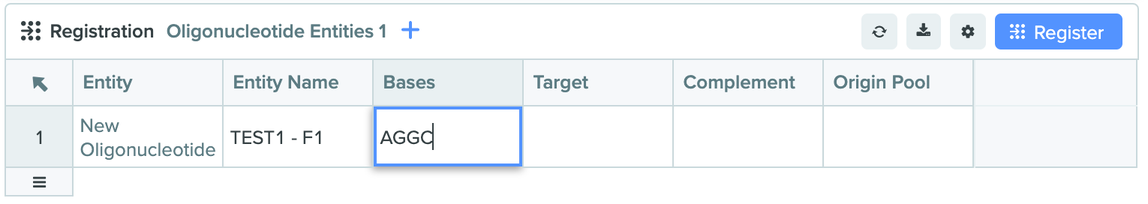
Specifying bases in Registration table for oligo entities: Registration tables for Oligo schema types now include a new column for recording sequences. This update consolidates the two step process of registering oligo entities and then updating base sequences of those entities to a single step within a Notebook entry. This saves scientists’ time and improves ease of use.
 Molecular Biology
Molecular Biology
-
Analyze translations in forward and reverse orientations: With the latest update, scientists can now create translations in either the forward or the reverse directions with ease. Additionally, the biochemical properties of these translations can be easily compared by toggling between the two analyses. This enhancement significantly speeds up analysis time and delivers relevant calculations with just a few clicks.
-
Primer organization and fields: To make working with Primers easier and more intuitive, we released several updates. These changes enable the selection of subsets of primers and streamline the use of primers in your experiment. Added checkboxes in the primers menu allow selection of a subset of attached primers.“Fill Primer” field option added as a schema field. This provides the option to fill-in selected primers in appropriate fields in the sequence metadata.Moved “Link Primers” option from the primers tab into the sequence map context menu.
-
Repeat Bulk Assembly runs: The Molecular Biology app now supports the option to repeat the preceding Bulk Assembly run. This is helpful when you have to execute a lot of bulk runs with similar settings in a single session. This option disappears once you exit the Bulk Assembly tool. Learn more about our Bulk Assembly tool here.
-
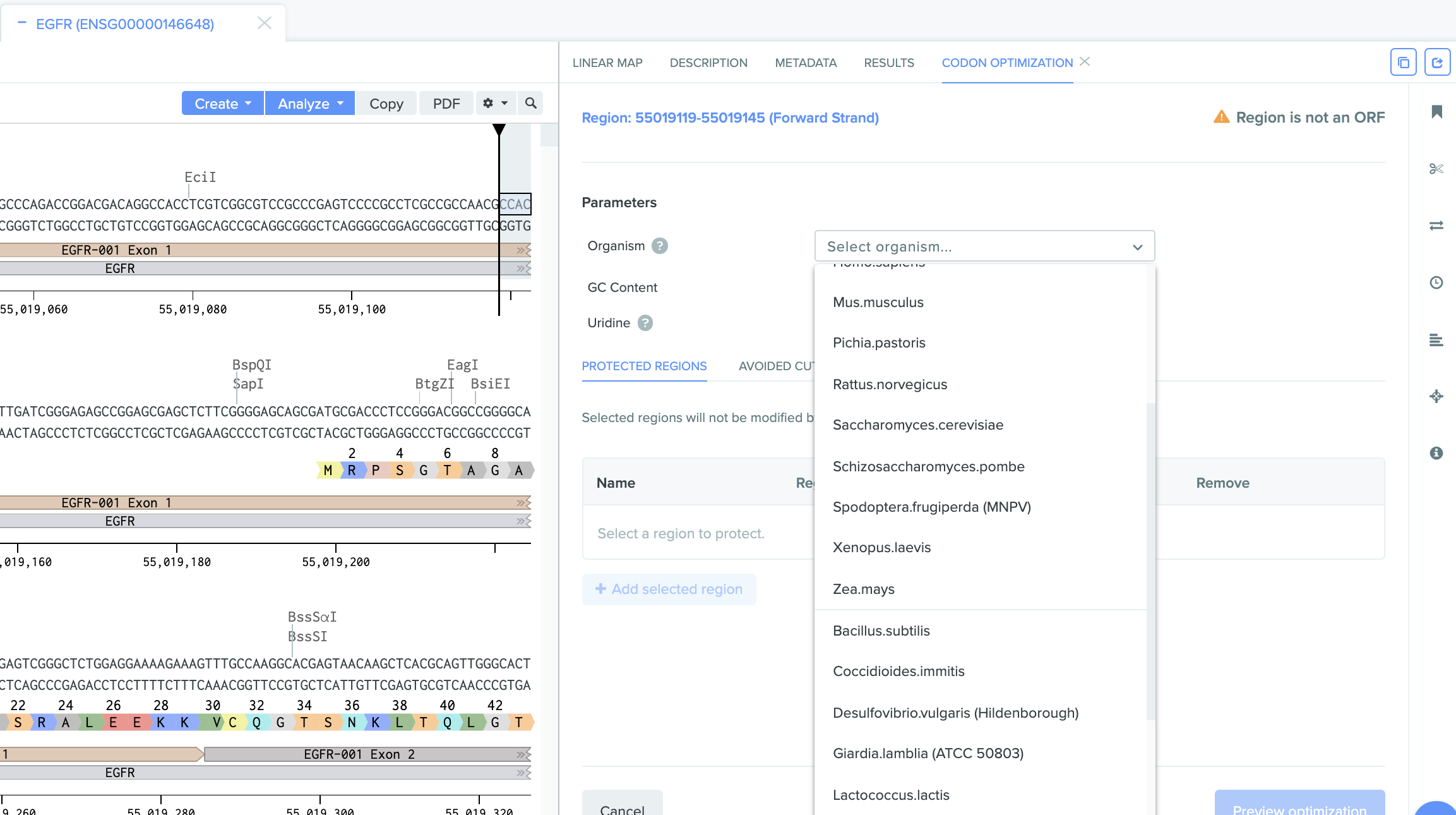
Added more organism codon usage tables for codon optimization: We continue to add more organisms to the codon optimization tool to better support emerging scientific needs. Scientists can now optimize the expression of proteins in 29 different host organisms on Benchling. This vastly expands our ability to support diverse codon optimization research needs.
Registry
Registry is core to modeling data on Benchling and managing all data related to critical samples. However, as R&D needs change, it is important for the Registry to evolve accordingly. We released features which make it easy to archive historical data and significantly simplify oligo creation.
-
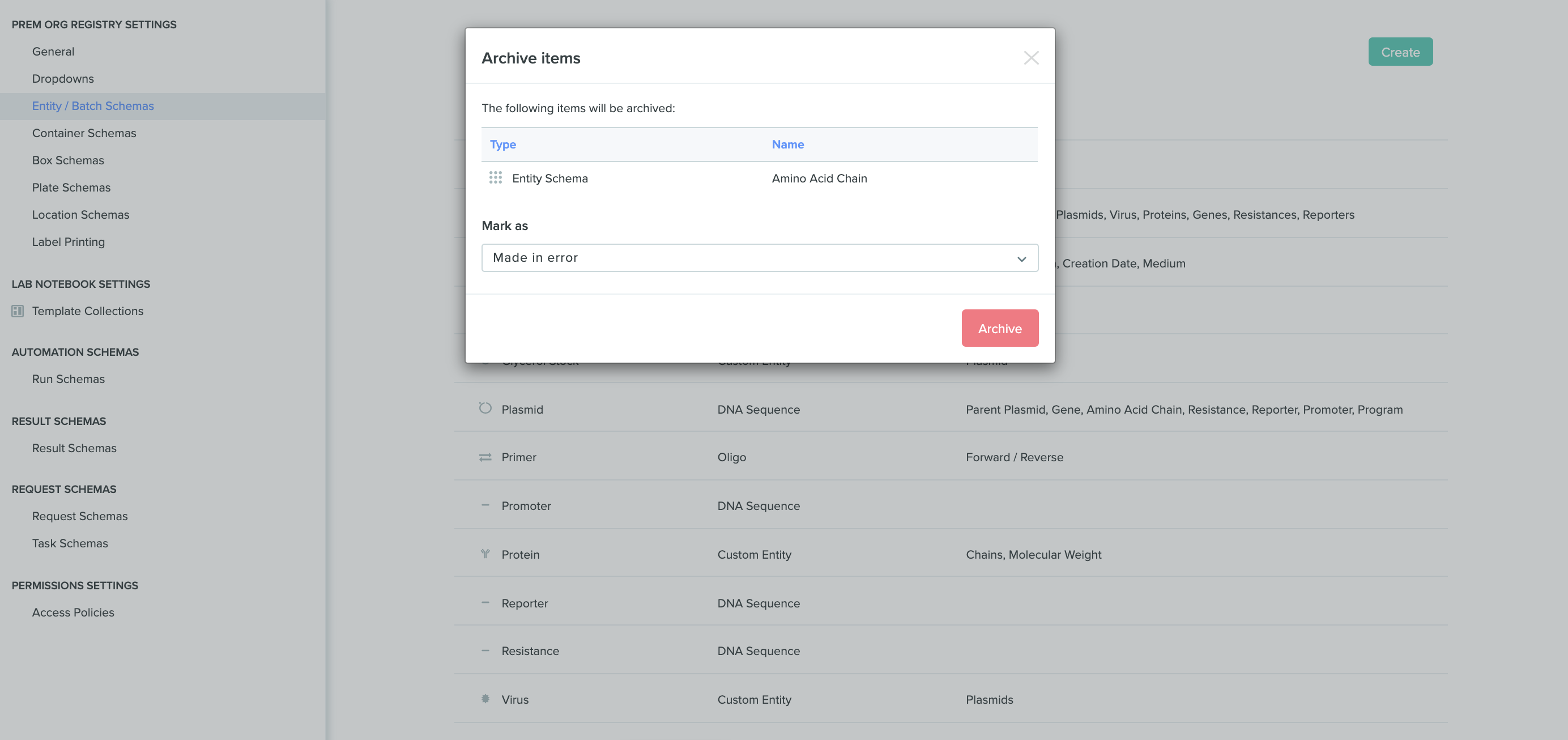
Archive a Registry schema without archiving its existing entities: Schemas can now be archived without the need to delete all existing entities that are using that schema. This helps preserve historic data while maintaining flexibility to create new data models. To archive a schema, navigate to the registry settings, hover over the schema you wish to archive, click on the red Archive button, and select the reason for archival.
-
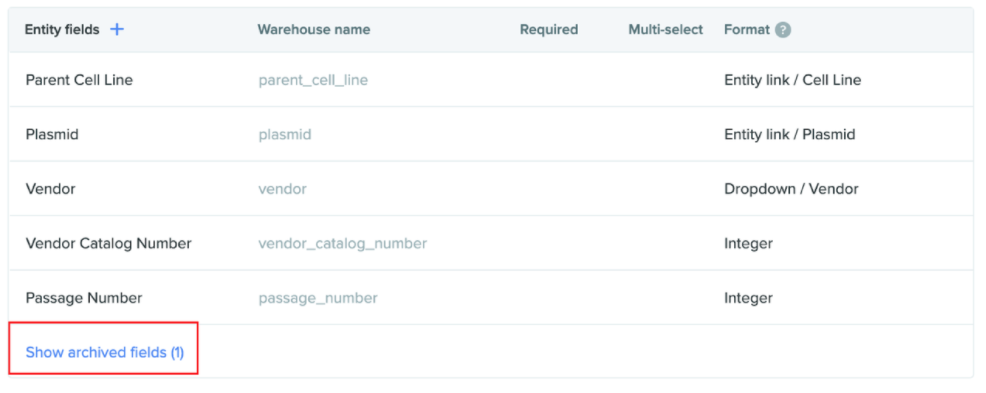
Hide/show archived schema fields: All schema configuration pages now have the option to either show or hide archived fields. All schemas with archived fields will have this feature. The archived fields will be set to hide by default. This provides easy access to all schema fields either current or old in one place.
-
Input bases during oligo entity creation: Oligo creation is now easier than ever in Benchling. Within the Registry, there is now a new field to input bases directly during Oligo entity creation. This combines two discrete steps in one simple step, saving scientists time and effort.
 Inventory
Inventory
-
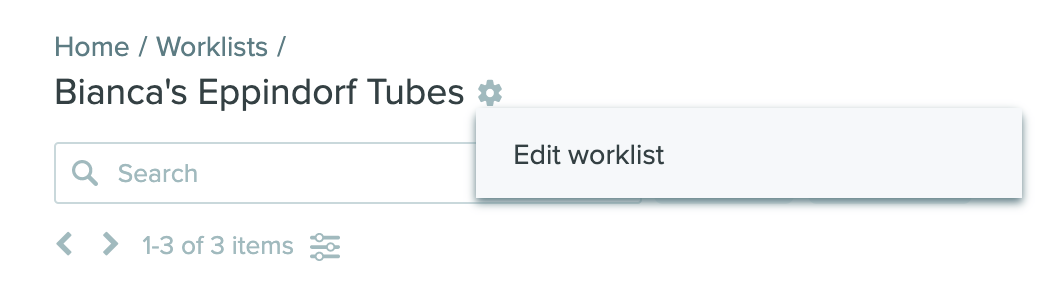
Renaming Worklists: Users can now rename Worklists directly in the UI without the need to delete in order to rename. Select the gear icon next to the Worklist name, then click edit Worklist. This update makes it easy to use and re-use existing Worklists to meet changing project needs.
-
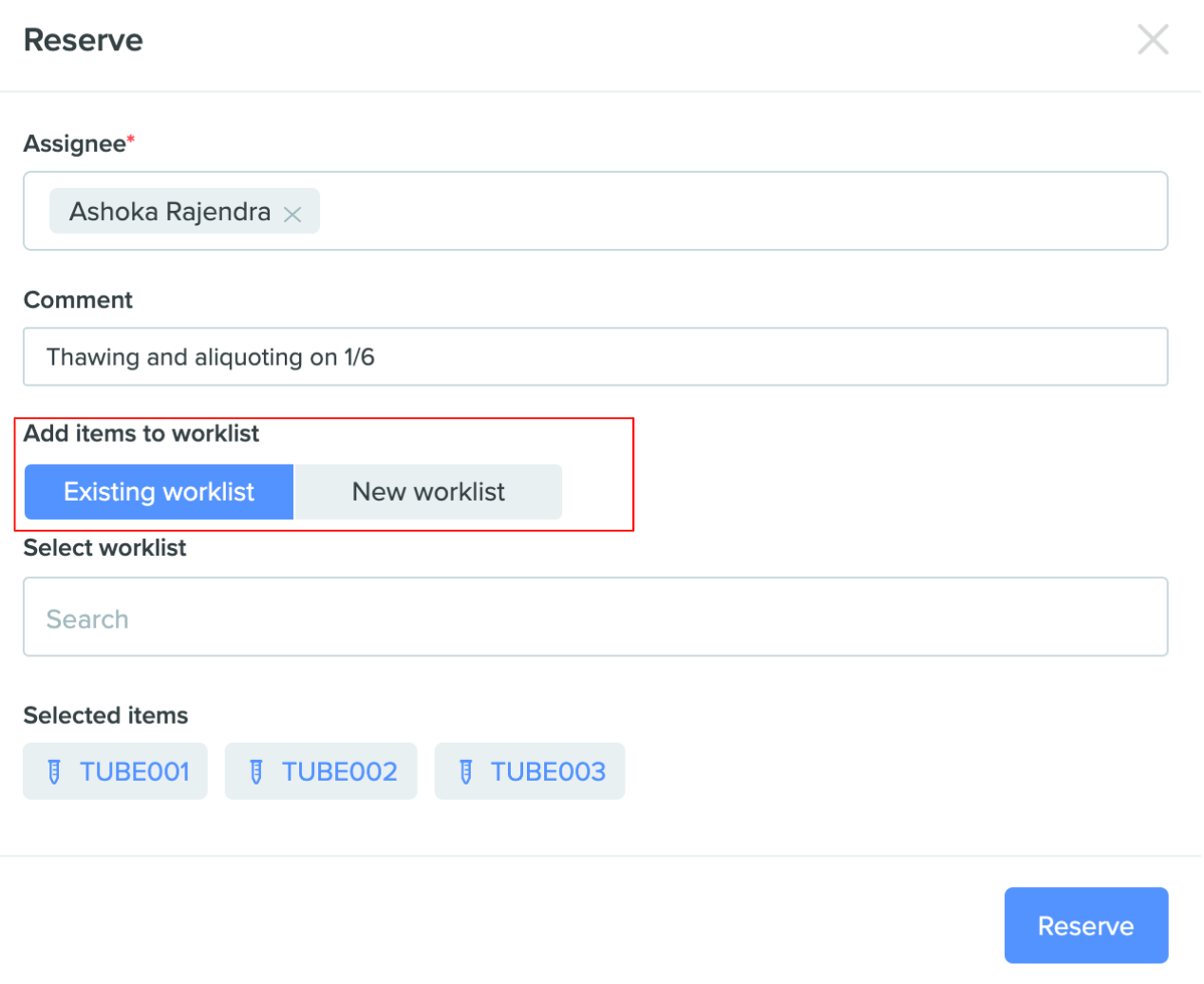
Create new Worklist when checking out or reserving inventory items: With this new update it is easy to either create a Worklist or add items to an existing worklist when checking out or reserving containers. The new UI modal allows users to quickly toggle between the two options. This makes the Worklist feature more intuitive to use along with the check out / reservation feature and better maps actual use cases in the lab.
-
Notification while archiving checked out or reserved containers: We added a notification when attempting to archive checked-out or reserved containers. Before proceeding to archive containers, the reserved / checked-out status will be displayed in orange text next to the selected container. This helps teams better manage their inventory items and mitigates possible erroneous actions on Benchling.
 Requests
Requests
-
Request creation on behalf of a user: A Request can be created on behalf of another user by editing the “Requested By” field when the request is created. By default, the user creating the request is shown here, but the field can be updated to a different user. Once a Request has been created, subsequent updates to the “Requested by” field can only be done via the API. This feature adds more flexibility to teams of users that are migrating historical data to Benchling or have more sophisticated Request submission processes.
 App Platform and Admin
App Platform and Admin
-
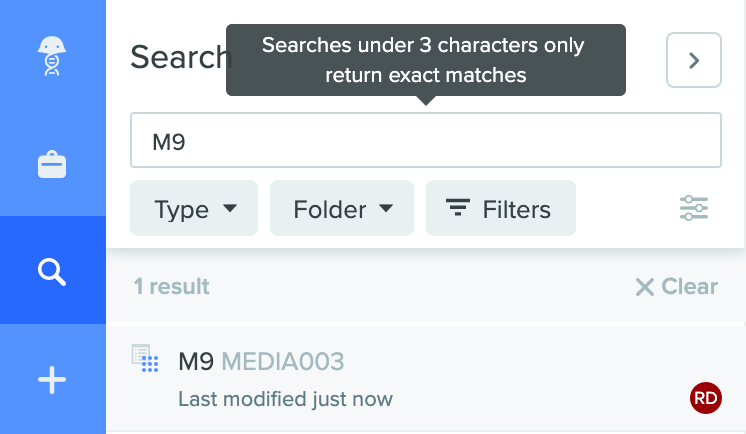
Search with 1+ character names: The global search tool on the main navigational toolbar now accepts searching for names that consist of 1 character or more. Please note that searches under 3 characters will only return exact matches. This update improves the utility of our search engine and helps find specific entities with short names faster.
-
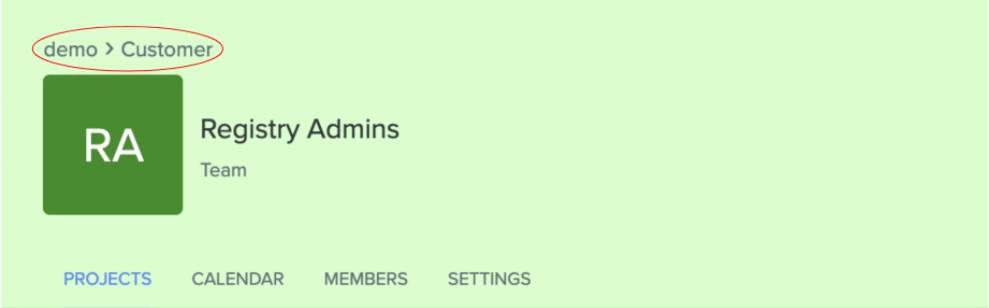
Improved navigation to the Tenant Admin Console: Tenant administrators can now more easily navigate back to the Tenant Admin Console from the org-dashboard or team-dashboard page via breadcrumb navigation at the top of the page. This improves accessibility of the dashboard and makes it easy for org admins to manage their users.
-
Increased number of users displayed in the Tenant Admin Console: To increase the ease of managing a large number of users, the user page in the Tenant Admin Console now displays up to 100 users. This is an increase from the previous display of 50 users. Note that this display number is not user-configurable.
Developer Platform
Our developer platform is fundamental to centralizing and standardizing all your R&D data. Benchling’s APIs are built to match the flexibility and speed of modern life science R&D. Keeping with this philosophy we released several API updates which super charge your research.
-
Update Run schema fields using the API: Benchling now supports updating Run schema fields by using the API via “PATCH /assay-runs/<run_id>”. The run_id refers to the ID of the run that will be updated. This endpoint takes in fields, if specified, and replaces those field values with new values. Learn more about runs here.
-
Search for sequences by exact base match: Benchling’s API now has a new search filter that allows searching for sequences with exact base match.
-
Attach primers to DNA sequences: Benchling now supports attaching primers (oligo ID or bases) to DNA sequences using API.
For a more comprehensive list of all the features released refer to the technical release notes here.